|
|
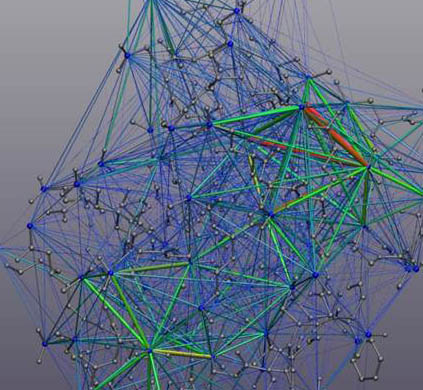
Multi-scale 4-D analytics for MD
trajectories.In support of two new papers we released in version 1.5 an improved BADE solver for mutal
information. We also extended TimeScapes to lipids and solvent analysis
(this code will be released shortly, for inquiries please
write to the e-mail address below).
Version 1.4: Advanced Nonlinear Statistics using Multual Information. In support of a new paper (Kovacs and Wriggers, 2016), we have implemented a new fast information matching (FIM) solver that bridges
between fast and slow time scales and generates 3-D spatial heat map images
from a statistical analysis of 1-D MD trajectory time series.
TimeScapes is
a Python-based package that can be used to detect
and characterize significant conformational changes in simulated
biomolecular systems. The program makes use of multiple time scales of dynamics and spatial organization.
|
|
|